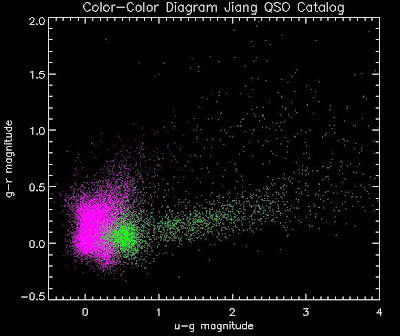
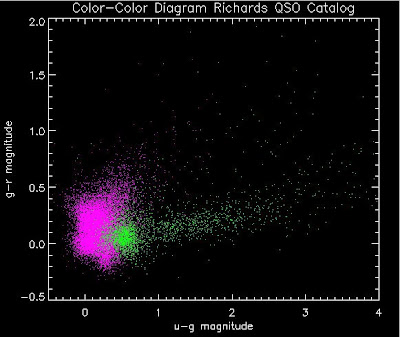
thisfile = './qsosMergedInfo.dat'
writecol,thisfile, qsos.ra, qsos.dec, qsos.z, qsos.zwarning, $
qsos.flux_clip_mean[0],qsos.flux_clip_mean[1], qsos.flux_clip_mean[2], $
qsos.flux_clip_mean[3], qsos.flux_clip_mean[4], fmt='(f,f,f,f,f,f,f,f,f)'
SDSS * Galaxy Clustering * Énergie Noire * Quasars

from pylab import *
from correlationFunctions import *
#------------------------------------------------------------------------
# Create file names (tiny catalogs)
#------------------------------------------------------------------------
workingDir = 'tinyrun'
makeworkingdir(workingDir)
galaxyDataFile, qsoDataFile, randomDataFile, corr2dCodefile, argumentFile, runConstantsFile = makeFileNamesTiny(workingDir)
oversample = 5. # Amount that randoms should be oversampled
corrBins = 25.0 # Number of correlation bins (+1)
mincorr = 0.1 # (Mpc/h comoving distance separation) Must be great than zero if log-binning
maxcorr = 10.0 # (Mphc/h comoving distance separation)
convo = 180./pi # conversion from degrees to radians
tlogbin = 1 # = 0 for uniform spacing, = 1 for log spacing in theta
#------------------------------------------------------------------------
# Write run constants to a file
#------------------------------------------------------------------------
writeRunConstantsToFile(runConstantsFile, galaxyDataFile, qsoDataFile, \
randomDataFile, corr2dCodefile, argumentFile, oversample, corrBins, \
mincorr, maxcorr, tlogbin)
#------------------------------------------------------------------------
# Compute the Angular Correlation Function
#------------------------------------------------------------------------
runcrossCorrelation(workingDir, argumentFile, corr2dCodefile, galaxyDataFile,\
qsoDataFile, randomDataFile, mincorr, maxcorr, corrBins, tlogbin)
# separation (Mpc/h) crossw (Mpc/h)
0.4300000000 -0.1156862745
1.0900000000 -0.1044776119
1.7500000000 -0.1208039566
2.4100000000 -0.0914845135
3.0700000000 -0.0393970538
3.7300000000 -0.0268417043
4.3900000000 0.0134841235
5.0500000000 0.0596093513
5.7100000000 0.0227161938
6.3700000000 0.1025539385
7.0300000000 0.0929232804
7.6900000000 0.0900670231
8.3500000000 0.0591397849
9.0100000000 0.0284723490
9.6700000000 0.0598689436
As you can see the correlation functions match!




θ = 2πu
φ = cos-1(2v - 1)
dΩ = sinφdθdφ = -dθd(cosφ)
randomsize = 100
randomtheta = findgen(randomsize)*0.0
randomphi = findgen(randomsize)*0
index = 0
while index lt randomsize do begin
thisindex = index
while index eq thisindex do begin
phi = 2*!pi*RANDOMU(S, 1)
theta = acos(2*RANDOMU(S, 1) - 1)
ang2pix_nest, nside_in, theta, phi, ipnest
thisweight = weight[ipnest]
if thisweight gt 2.0 then begin
randomtheta[index] = theta
randomphi[index] = phi
index = index+1
endif
endwhile
endwhile