Wednesday, January 11, 2012
119th AAS Meeting, Austin Texas

Thursday, November 3, 2011
Talk at Brookhaven
My hosts Erin Sheldon and Anže Slosar gave me some really helpful feed back:
- I wasn't expecting such a mixed audience, and I didn't give enough introduction to BOSS, and the data. For instance I didn't directly discuss why quasars are hard to target. I also didn't emphasize that the inputs to the likelihood were the photometric fluxes. I should have talked more about the difference between photometric and spectroscopic data. But in general, I need to be more aware of the audience and beef up the introduction.
- I went through the full derivation of the likelihood, and this wasn't necessary.
- I was nervous and relied on notecards. Anže said this made it look like I didn't understand what I was talking about. Basically, I need to practice the talk a bunch so I don't need cards.
- I mispoke and talked about the first BAO detection. I said it was from SDSS-II, it was actually from SDSS-I.
- In general I didn't feel prepared to answer all questions, I need to go through the talk and make sure I fully understand everything I am posting -- for instance how was the first BAO detection made?
Wednesday, September 7, 2011
More streamlining
Black Hole Masses
Saturday, August 13, 2011
Tuesday, June 14, 2011
Temperature Plots in Python
import matplotlib
import numpy as np
import matplotlib.cm as cm
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
redbins = linspace(start = 0.5, stop = 1.0, num = 51)
rcenter = (redbins[0:-1]+redbins[1:])/2.0
mbins = linspace(start = -28, stop = -18, num = 51)
mcenter = (mbins[0:-1]+mbins[1:])/2.0
H, xedges,yedges = N.histogram2d(qsoM[dr7cut],qsored[dr7cut],bins=(mbins,redbins))
X = rcenter
Y = mcenter
Z = H
#plt.imshow(H, extent=[-3,3,-3,3], interpolation='bilinear', origin='lower')
#plt.colorbar()
plt.imshow(H, extent=[-3,3,-3,3], interpolation='nearest', origin='lower')
plt.colorbar()
xlabel('redshift (z)')
ylabel('absolute magnitude (i-band)')
title('SDSS DR7 Quasar Density')
The above code is also in the following log file: ../logs/110614log.py

Thursday, June 9, 2011
More QSO-Galaxy Cross Correlations
Monday, June 6, 2011
Testing QSO-Galaxy Cross CF
from pylab import *
from correlationFunctions import *
#------------------------------------------------------------------------
# Create file names (tiny catalogs)
#------------------------------------------------------------------------
workingDir = 'tinyrun'
makeworkingdir(workingDir)
galaxyDataFile, qsoDataFile, randomDataFile, corr2dCodefile, argumentFile, runConstantsFile = makeFileNamesTiny(workingDir)
oversample = 5. # Amount that randoms should be oversampled
corrBins = 25.0 # Number of correlation bins (+1)
mincorr = 0.1 # (Mpc/h comoving distance separation) Must be great than zero if log-binning
maxcorr = 10.0 # (Mphc/h comoving distance separation)
convo = 180./pi # conversion from degrees to radians
tlogbin = 1 # = 0 for uniform spacing, = 1 for log spacing in theta
#------------------------------------------------------------------------
# Write run constants to a file
#------------------------------------------------------------------------
writeRunConstantsToFile(runConstantsFile, galaxyDataFile, qsoDataFile, \
randomDataFile, corr2dCodefile, argumentFile, oversample, corrBins, \
mincorr, maxcorr, tlogbin)
#------------------------------------------------------------------------
# Compute the Angular Correlation Function
#------------------------------------------------------------------------
runcrossCorrelation(workingDir, argumentFile, corr2dCodefile, galaxyDataFile,\
qsoDataFile, randomDataFile, mincorr, maxcorr, corrBins, tlogbin)
# separation (Mpc/h) crossw (Mpc/h)
0.4300000000 -0.1156862745
1.0900000000 -0.1044776119
1.7500000000 -0.1208039566
2.4100000000 -0.0914845135
3.0700000000 -0.0393970538
3.7300000000 -0.0268417043
4.3900000000 0.0134841235
5.0500000000 0.0596093513
5.7100000000 0.0227161938
6.3700000000 0.1025539385
7.0300000000 0.0929232804
7.6900000000 0.0900670231
8.3500000000 0.0591397849
9.0100000000 0.0284723490
9.6700000000 0.0598689436
As you can see the correlation functions match!
Wednesday, June 1, 2011
Data in Order
Tuesday, May 24, 2011
Data Handling
Alexie wrote a pipeline to collate data and apply masks. The code is here: .../qsobias/Analysis/doall_cfht_se.pro
I added some code to remove the repeats: ../qsobias/Analysis/remove_repeats.pro
To just start us off I wrote some code to trim the data and make randoms in the entire footprint (because we don't have masks yet). This code is in the following log file: .../logs/110522log.pro
;I start with the galaxy catalog from Alexie: ../Catalogs/cs82_all_cat6.fits
;I remove repeats:
cat6 = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cs82_all_cat6.fits'
cat7 = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cs82_all_cat7.fits'
infile = cat6
outfile = cat7
remove_repeats,infile,outfile
;Read in file with repeats removed
infile = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cs82_all_cat7.fits'
str=mrdfits(infile,1)
;Do some simple geometry cuts on ra, dec, and z
deltacut = where(str.DELTA_J2000 gt -1.0 and str.DELTA_J2000 lt 0.94)
str = str[deltacut]
alphacut = where(str.ALPHA_J2000 gt -42.5 and str.ALPHA_J2000 lt 45.0)
str = str[alphacut]
zcut = where(str.ZPHOT le 1.0)
str = str[zcut]
;Do some cuts that Alexie recommends to make sure we are getting galaxies, inside the mask, and only down to mag = 23.7
gooddatacut = where(str.mask eq 0 and str.mag_auto lt 23.7 and str.class_star lt 0.9)
str = str[gooddatacut]
;Outfile data to files
outfile = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cs82_all_cat8.fits'
mwrfits, str, outfile, /create
;This is the galaxy catalog I am using for the preliminary correlation function test
thisfile = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cfht_data.dat'
writecol,thisfile,str.alpha_j2000,str.delta_j2000,str.zphot
;Now to do the same to the BOSS Data
;This code can be found in 110519log.py
;Read in BOSS Galaxy Data
qsofile = '/home/jessica/qsobias/Stripe_82_QSOs_unique.dat'
readcol,qsofile,ra,dec,z,gmag,format='(D,D,D,D,X)'
;Remove duplicates
spherematch, ra, dec, ra, dec, 2./3600, m1, m2, maxmatch=0
dups = m1[where(m1 gt m2)]
good = [indgen(n_elements(ra))*0]+1
good[dups]=0
ra = ra[where(good)]
dec = dec[where(good)]
z = z[where(good)]
gmag = gmag[where(good)]
thisfile = '/home/jessica/qsobias/QSOs_unique.dat'
writecol,thisfile,ra,dec,z,gmag
#Now in python
import numpy as N
from pylab import *
from dataplay import *
galfile = '/clusterfs/riemann/data/jessica/alexieData/Catalogs/cfht_data.dat'
gal=N.loadtxt(galfile,comments='#')
galra = gal[:,0]
galdec = gal[:,1]
galz = gal[:,2]
#Make Cuts on QSOs
qsofile = '/home/jessica/qsobias/QSOs_unique.dat'
qso=N.loadtxt(qsofile,comments='#')
qsora = qso[:,0]
qsodec = qso[:,1]
qsoz = qso[:,2]
qsog = qso[:,3]
#Dec between -1 and 1
deccut = N.where((qsodec < 0.94) & (qsodec > -1.0)) #make sure they are inside the cfht mask
qsora = qsora[deccut]
qsodec = qsodec[deccut]
qsoz = qsoz[deccut]
qsog = qsog[deccut]
#Redshift between 0 and 1
zcut = N.where((qsoz >= 0.0) & (qsoz < 1.0)) qsora = qsora[zcut] qsodec = qsodec[zcut] qsoz = qsoz[zcut] qsog = qsog[zcut] #RA range (-42.5,45) highra = N.where(qsora > 300)
qsora[highra] = qsora[highra] - 360.
racut = N.where((qsora < 45.0) & (qsora > -42.5))
qsora = qsora[racut]
qsodec = qsodec[racut]
qsoz = qsoz[racut]
qsog = qsog[racut]
thisfile = '/home/jessica/qsobias/QSO_data_masked.dat'
writeQSOsToFile(qsora,qsodec,qsoz,qsog,thisfile)
thisfile = '/home/jessica/qsobias/QSO_data.dat'
writeQSOsToFile2(qsora,qsodec,qsoz,thisfile)
plot(galra, galdec, 'b,', label = 'galaxy data')
plot(qsora, qsodec, 'r,', label = 'qso data')
xlabel('ra')
ylabel('dec')
title('RA & Dec of Data')
from pylab import *
legend(loc=1)


Friday, May 20, 2011
New Project
Here is the new project description:
Participants:
Jessica Kirkpatrick
Martin White, David Schlegel, Nic Ross, Alexie Leauthaud, Jean-Paul Kneib
Categories: BOSS
Project Description:
We plan to measure the intermediate scale clustering of low redshift BOSS quasars along Stripe 82 by cross-correlation against the photometric galaxy catalog from the CFH i-band imaging on Stripe 82. There are approximately 1,000 BOSS quasars with 0.5<z<1 and just under 6 million galaxies in the -43<RA<43 and -1<DEC<1 region brighter than i=23.5, which should lead to a strong detection of clustering over approximately 2 orders of magnitude in length scale. The geometry of the stripe suggests errors on the cross-correlation can be efficiently obtained by jackknife or bootstrap sampling the ~50 2x2 degree blocks.
We intend to split the QSO sample in luminosity and black hole mass. We plan to estimate the BH mass using the fits from Vestergaard and Peterson, knowing the Hbeta line width and the continuum luminosity at 5100A. The pipeline measures the former, we plan to measure the latter from the photometry calibrated with Ian McGreer's mocks.
If we use the QG/QR-1 estimator we do not need the quasar mask, only that of the galaxies which will be provided by the CS82 team in the form of a pixelized mask from visual inspection. The dN/dz of the galaxies is known from photometric redshifts plus spectroscopic training sets. While the galaxies could be split in photometric redshift bins, the gains from doing so are not expected to be large, so our initial investigations will simply cross-correlate the quasars with the magnitude limited galaxy catalog.
Along with this project we will submit a request for EC status for:
Ludo van Waerbeke
Hendrik Hildebrant
David Woods
Thomas Erben
who were instrumental in obtaining and reducing the CS82 data and producing the required galaxy catalog and mask but are not members of the BOSS collaboration.
Details are available at https://www.sdss3.org/
The first thing Martin wanted me to do was to approximate the errors bars for the correlation function based on the density of galaxies and quasars in my sample.
Starting with luminosity function in this paper, I am doing the following to estimate the density.
According to Table 1 in Ilbert et. al. the following are the Schechter parameters for the galaxy luminosity function, redshift 0.6-0.8:

I understand that the errors go as 1 / sqrt(pair counts) in each bin. But going from galaxy/qso density to pair counts in a bin is where I am a bit lost.
For a 3D correlation function the number of data pairs goes as Nqso times Nbar-galaxy times 1+xi times the volume of the bin (in 3D, e.g. 4\pi s^2 ds for a spherical shell). Just think of what the code does: sit on each quasar and count all the galaxies in the bin. To go from a 3D correlation function to a 2D correlation function you need to integrate in the Z direction. But remember that the sum of independent Poisson distributions is also a Poisson with a mean equal to the sum of the means of the contributing parts. So this allows you to figure out what the error on wp is. You should see that as you integrate to very large line-of-sight distance things become noisier. So choose something like +/-50Mpc/h for the width in line-of-sight distance to integrate over in defining wp.
It's a little easier to understand if you write the defining equations out for yourself on a piece of paper.
Martin
http://arxiv.org/abs/0802.2105
https://trac.sdss3.org/wiki/BOSS/quasars/black_hole_masses
Sunday, March 13, 2011
Unique Quasars
I wasn't exactly sure how to define "unique" quasar.
The simulated i-band fluxes range from 15 to 22.5. They are simulated to 4 decimal places.
The simulated redshifts (z) range from 0.5 to 5.0. They are simulated out to 6 decimal places.
If I define unique to be the same out to the maximum decimal places provided, then there are only 2 simulated quasars that are repeats of each other (0.00002%)
If I define unique to be within 0.1% magnitude/redshift of each other, then there are 6285 (out of 10 million) are repeats (0.06%).
If I define unique to be within 1% magnitude/redshift of each other, then there are 62,824 repeat objects (0.6%).
Anyway, so I guess I should ask Joe how he wants me to define repeats.
Here is the code to do this:
file = '/clusterfs/riemann/raid001/jessica/boss/Likeli_QSO.fits.gz'
qsos = mrdfits(file, 1)
imean = mean(qsos.i_sim)
zmean = mean(qsos.z_sim)
percent = 1.0/100. #simulated quasars within this percentage of each other
ierr = imean*percent
zerr = zmean*percent
imult = round(1./ierr)*1.
zmult = round(1./zerr)*1.
roundqsosz = round(qsos.z_sim*zmult)/zmult
roundqsosi = round(qsos.i_sim*zmult)/imult
uniqz = uniq(roundqsosz)
uniqi = uniq(roundqsosi)
all = where(qsos.z_sim GT -99999)
repeatZ = setdifference(all, uniqz)
repeatI = setdifference(all, uniqI)
help, setintersection(repeatZ, repeatI)
Also can be found in the following log file:
../logs/110313log.pro
Thursday, May 20, 2010
New Threshold Results
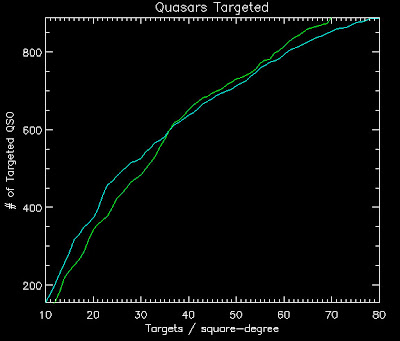
Below are the lratio new thresholds for the old version of the likelihood (v1) and the likelihood with ALL the Quasars as a function of targets per square degree (TPSD):
TPSD ----- v1 threshold - all quasar threshold
10.0000 -----0.749085 --------- 0.770306
20.0000 -----0.490932 --------- 0.571854
30.0000 -----0.347334 --------- 0.425242
40.0000 -----0.253020 --------- 0.329808
50.0000 -----0.198630 --------- 0.264506
60.0000 -----0.160074 --------- 0.214092
70.0000 -----0.128912 --------- 0.182231
80.0000 -----0.107913 --------- 0.152691
Cyan is the ALL the Quasars
Green is likelihood v1

Using these thresholds here are the number of quasars targeted:
TPSD ---- #QSOs (v1) --- #QSOs (ALL Quasars)
10.0000 ---443.000 -------- 406.000
20.0000 ---656.000 -------- 582.000
30.0000 ---781.000 -------- 722.000
40.0000 ---897.000 -------- 830.000
50.0000 ---965.000 -------- 892.000
60.0000 ---1035.00 -------- 955.000
70.0000 ---1089.00 -------- 1000.00
80.0000 ---1123.00 -------- 1037.00
Cyan is the ALL the Quasars
Green is likelihood v1

Looks like ALL the quasars doesn't do better after all with the non-Milky Way thresholds. Boo.
The log file to run this code is here: ../logs100520log.pro
Monday, May 17, 2010
ALL the Quasars Results
Below are the lratio thresholds for the old version of the likelihood (v1) and the likelihood with ALL the Quasars as a function of targets per square degree (TPSD):
TPSD ----- v1 threshold - all quasar threshold
10.0000 ----- 0.996883 ----- 0.984691
20.0000 ----- 0.868663 ----- 0.795943
30.0000 ----- 0.674674 ----- 0.629976
40.0000 ----- 0.496851 ----- 0.522806
50.0000 ----- 0.403819 ----- 0.441149
60.0000 ----- 0.318171 ----- 0.363089
70.0000 ----- 0.262071 ----- 0.310237
80.0000 ----- 0.215056 ----- 0.268664
Cyan is the ALL the Quasars
Green is likelihood v1
Using these thresholds here are the number of quasars targeted:
TPSD ---- #QSOs (v1) --- #QSOs (ALL quasars)
10.0000 -- 132.000 ----- 155.000
20.0000 -- 343.000 ----- 374.000
30.0000 -- 484.000 ----- 527.000
40.0000 -- 651.000 ----- 638.000
50.0000 -- 731.000 ----- 713.000
60.0000 -- 815.000 ----- 793.000
70.0000 -- 888.000 ----- 853.000
80.0000 -- 944.000 ----- 889.000
Cyan is the ALL the Quasars
Green is likelihood v1
Wednesday, May 12, 2010
ALL the Quasars
Before we had a brightness limit and some other cuts, but Hennawi wanted me to see what it looked like with all of them.
I did this by modifying: qso_fake_jess.pro with the following call to qso_photosamp:
qsos = qso_photosamp(Z_MIN = Z_MIN, Z_MAX = Z_MAX, NOCUTS = NOCUTS, ILIM = 23.0)
.run hiz_kde_numerator_jess.pro
To create a new QSO Catalog:
../likelihood/qsocatalog/QSOCatalog-Wed-May-12-14:03:41-2010.fits
; Read in the QSO file
file2 = '/home/jessica/repository/ccpzcalib/Jessica/likelihood/qsocatalog/QSOCatalog-Wed-May-12-14:03:41-2010.fits'
This run is in the directory:
../likelihood/likev1all
The log file is here:
../logs/100512log.pro
~~~~~~~~
this, leave a comment so I know you are out there!
~~~~~~~~
Tuesday, May 11, 2010
Likelihood Threshold Results
Below are the thresholds for the old version of the likelihood (v1) in parentheses are the thresholds Adam Myers got:
Targets/deg2---------Threshold
10----------------0.995851
20----------------0.858535 (0.533)
30----------------0.664424
40----------------0.490932 (0.235)
50----------------0.400611
60----------------0.311577
70----------------0.256615
80----------------0.213374
So my thresholds still don't match his.
In terms of recovered QSOs with these thresholds:
Targets/deg2---Threshold-----Recovered QSOs (out of 1625)
10.0000 --- 0.995851 ------- 135 (8.3%)
20.0000 --- 0.858535 ------- 346 (21.3%)
30.0000 --- 0.664424 ------- 489 (30.1%)
40.0000 --- 0.490932 ------- 651 (40.1%)
50.0000 --- 0.400611 ------- 727 (44.7%)
60.0000 --- 0.311577 ------- 816 (50.2%)
70.0000 --- 0.256615 ------- 886 (54.5%)
80.0000 --- 0.213374 ------- 942 (58.0%)
I've saved these likelihoods in the directory: ../likelihood/likev1/
The they are also here:
likelihoodv1thresholds.fits
qsolikelihoodv1thresholds.fits
The log file is here: ../logs/100511log.pro
Now to make some magic happen...
Tuesday, April 27, 2010
Likelihood Test Results (4)
eps = 1e-30
bossqsolike = total(likelihood.L_QSO_Z[16:32],1) ;quasars between redshift 2.1 and 3.7
qsolcut1 = where( alog10(total(likelihood.L_QSO_Z[16:32],1)) LT -9.0)
den = total(targets.L_EVERYTHING_ARRAY[0:4],1) + total(likelihood.L_QSO_Z[0:44],1) + eps
num = bossqsolike + eps
NEWRATIO = num/den
NEWRATIO[qsolcut1] = 0 ; eliminate objects with low L_QSO value
IDL> print, n_elements(ql) ; number quasars
461
IDL> print, n_elements(ql)*1.0/(n_elements(sl)+n_elements(ql)) ;percent accuracy
0.256111
IDL> print, n_elements(ql) + n_elements(sl) ; total targeted
1800
IDL>
IDL> print, n_elements(nql) ; number quasars
575
IDL> print, n_elements(nql)*1.0/(n_elements(nsl)+n_elements(nql)) ;percent accuracy
0.319444
IDL> print, n_elements(nql) + n_elements(nsl) ; total targeted
We now go from 25.6% accuracy to 31.9% accuracy! And we find 114 more quasars!
Monday, April 26, 2010
Likelihood Test Results (3)
It seems that adding in the BOSS QSOs doesn't help.
Now I am going to run on the old qso catalog but with the larger redshift range (0.5 < z < 5.0) and see if this improves things. See ../logs/100426log.pro for code.
So this seems to actually make a difference. I have played around with the redshift range for the numerator to get the best selection numbers:
eps = 1e-30
bossqsolike = total(likelihood.L_QSO_Z[14:34],1) ;quasars between redshift 2.1 and 3.5
qsolcut1 = where( alog10(total(likelihood.L_QSO_Z[19:34],1)) LT -9.0)
den = total(targets.L_EVERYTHING_ARRAY[0:4],1) + total(likelihood.L_QSO_Z[0:44],1) + eps
num = bossqsolike + eps
NEWRATIO = num/den
NEWRATIO[qsolcut1] = 0 ; eliminate objects with low L_QSO value
qsolcut2 = where( alog10(total(targets.L_QSO_Z[0:18],1)) LT -9.0)
L_QSO = total(targets.L_QSO_Z[2:18],1)
den = total(targets.L_EVERYTHING_ARRAY[0:4],1) + total(targets.L_QSO_Z[0:18],1) + eps
num = L_QSO + eps
OLDRATIO = num/den
OLDRATIO[qsolcut2] = 0 ; eliminate objects with low L_QSO value
numsel = 1800-1
NRend = n_elements(newratio)-1
sortNR = reverse(sort(newratio))
targetNR = sortNR[0:numsel]
restNR = sortNR[numsel+1:NRend]
ORend = n_elements(oldratio)-1
sortOR = reverse(sort(oldratio))
targetOR = sortOR[0:numsel]
restOR = sortOR[numsel+1:ORend]
nql = setintersection(quasarindex,targetNR)
nqnl = setintersection(quasarindex, restNR)
nsl = setdifference(targetNR, quasarindex)
nsnl = setdifference(restNR, quasarindex)
ql = setintersection(quasarindex, targetOR)
qnl = setintersection(quasarindex, restOR)
sl = setdifference(targetOR,quasarindex)
snl = setdifference(restOR,quasarindex)
IDL> print, n_elements(ql) ; number quasars
461
IDL> print, n_elements(ql)*1.0/(n_elements(sl)+n_elements(ql)) ;percent accuracy
0.256111
IDL> print, n_elements(ql) + n_elements(sl) ; total targeted
1800
IDL>
IDL> print, n_elements(nql) ; number quasars
508
IDL> print, n_elements(nql)*1.0/(n_elements(nsl)+n_elements(nql)) ;percent accuracy
0.282222
IDL> print, n_elements(nql) + n_elements(nsl) ; total targeted
1800
So we go from 25.6% accuracy to 28.2% accuracy! And we find 47 more quasars!
The white points were targeted/missed by both the new and old likelihoods
The magenta points were only targeted/missed by the new likelihood
The cyan points were only targeted/missed by the old likelihood

Now to see how the different luminosity functions do. Below is results running with the Richard 06 luminosity function:
(See ../logs/100427_2log.pro for code)
IDL> print, n_elements(ql) ; number quasars
461
IDL> print, n_elements(ql)*1.0/(n_elements(sl)+n_elements(ql)) ;percent accuracy
0.256111
IDL> print, n_elements(ql) + n_elements(sl) ; total targeted
1800
IDL>
IDL> print, n_elements(nql) ; number quasars
543
IDL> print, n_elements(nql)*1.0/(n_elements(nsl)+n_elements(nql)) ;percent accuracy
0.301667
IDL> print, n_elements(nql) + n_elements(nsl) ; total targeted
1800
This does better than the old luminosity function!
So we go from 25.6% accuracy to 30.2% accuracy! And we find 82 more quasars!
Sunday, April 25, 2010
Likelihood Test Results (2)
Joe suggested doing this. We'll see if it improves things, if not, then looks like we should go back to old catalog and see if modifying the luminosity function helps.
Here is what they look like:
The code to do this is in the following log file: ../logs/100425log.pro
It creates the following file to use as input to Monte Carlo: ~/boss/allQSOMCInput.fits"
../likelihood/qsocatalog/QSOCatalog-Mon-Apr-26-13:22:43-2010.fits
I changed likelihood_compute to look at the above catalog.
Run the likelihood calculation with this new input catalog:
smalltargetfile = "./smalltarget.fits"
smalltargets = mrdfits(smalltargetfile, 1)
.com splittargets2.pro
nsplit = 50L
splitsize = floor(n_elements(smalltargets)*1.0/nsplit)
junk = splitTargets2(smalltargets, nsplit, splitsize)
; run likelihood.script
.com mergeLikelihoods.pro
likelihood = mergelikelihoods(nsplit)
outfile = "./likelihoods2.0-4.0ALLBOSSAddedSmall.fits"
splog, 'Writing file ', outfile
mwrfits, likelihood, outfile, /create
IDL> print, n_elements(ql) ; number quasars
467
IDL> print, n_elements(ql)*1.0/(n_elements(sl)+n_elements(ql)) ;percent accuracy
0.259300
IDL> print, n_elements(ql) + n_elements(sl) ; total targeted
1801
IDL>
IDL> print, n_elements(nql) ; number quasars
418
IDL> print, n_elements(nql)*1.0/(n_elements(nsl)+n_elements(nql)) ;percent accuracy
0.232093
IDL> print, n_elements(nql) + n_elements(nsl) ; total targeted
1801
This doesn't do better than the old likelihood either (same color scheme as last post).
The logfile for this is ../logs/100425log.pro