thisfile = './qsosMergedInfo.dat'
writecol,thisfile, qsos.ra, qsos.dec, qsos.z, qsos.zwarning, $
qsos.flux_clip_mean[0],qsos.flux_clip_mean[1], qsos.flux_clip_mean[2], $
qsos.flux_clip_mean[3], qsos.flux_clip_mean[4], fmt='(f,f,f,f,f,f,f,f,f)'
SDSS * Galaxy Clustering * Énergie Noire * Quasars
from pylab import *
from correlationFunctions import *
#------------------------------------------------------------------------
# Create file names (tiny catalogs)
#------------------------------------------------------------------------
workingDir = 'tinyrun'
makeworkingdir(workingDir)
galaxyDataFile, qsoDataFile, randomDataFile, corr2dCodefile, argumentFile, runConstantsFile = makeFileNamesTiny(workingDir)
oversample = 5. # Amount that randoms should be oversampled
corrBins = 25.0 # Number of correlation bins (+1)
mincorr = 0.1 # (Mpc/h comoving distance separation) Must be great than zero if log-binning
maxcorr = 10.0 # (Mphc/h comoving distance separation)
convo = 180./pi # conversion from degrees to radians
tlogbin = 1 # = 0 for uniform spacing, = 1 for log spacing in theta
#------------------------------------------------------------------------
# Write run constants to a file
#------------------------------------------------------------------------
writeRunConstantsToFile(runConstantsFile, galaxyDataFile, qsoDataFile, \
randomDataFile, corr2dCodefile, argumentFile, oversample, corrBins, \
mincorr, maxcorr, tlogbin)
#------------------------------------------------------------------------
# Compute the Angular Correlation Function
#------------------------------------------------------------------------
runcrossCorrelation(workingDir, argumentFile, corr2dCodefile, galaxyDataFile,\
qsoDataFile, randomDataFile, mincorr, maxcorr, corrBins, tlogbin)
# separation (Mpc/h) crossw (Mpc/h)
0.4300000000 -0.1156862745
1.0900000000 -0.1044776119
1.7500000000 -0.1208039566
2.4100000000 -0.0914845135
3.0700000000 -0.0393970538
3.7300000000 -0.0268417043
4.3900000000 0.0134841235
5.0500000000 0.0596093513
5.7100000000 0.0227161938
6.3700000000 0.1025539385
7.0300000000 0.0929232804
7.6900000000 0.0900670231
8.3500000000 0.0591397849
9.0100000000 0.0284723490
9.6700000000 0.0598689436
As you can see the correlation functions match!




PRO t_histogramIf you add maxmatch=1 to your spherematch it will avoid duplicates (should default to this)
data = [[-5, 4, 2, -8, 1], $
[ 3, 0, 5, -5, 1], $
[ 6, -7, 4, -4, -8], $
[-1, -5, -14, 2, 1]]
hist = HISTOGRAM(data)
bins = FINDGEN(N_ELEMENTS(hist)) + MIN(data)
PRINT, MIN(hist)
PRINT, bins
SPLOT, bins, hist, YRANGE = [MIN(hist)-1, MAX(hist)+1], PSYM = 10, XTITLE = 'Bin Number', YTITLE = 'Density per Bin'
END
/home/jessica/boss/BOSS_QSOs_First14plates.fitsTarget bitmasks from the wiki:
/home/jessica/boss/bosstarget-qso-comm-collate.fits
# restrictive qso selection
maskbits BOSS_TARGET1 10 QSO_CORE
# permissive qso selection
maskbits BOSS_TARGET1 11 QSO_BONUS
# known qso between [2.2,9.99]
maskbits BOSS_TARGET1 12 QSO_KNOWN_MIDZ
# known qso outside of miz range. never target
maskbits BOSS_TARGET1 13 QSO_KNOWN_LOHIZ
# Neural Net that match to sweeps/pass cuts
maskbits BOSS_TARGET1 14 QSO_NN
# UKIDSS stars that match sweeps/pass flag cuts
maskbits BOSS_TARGET1 15 QSO_UKIDSS
# kde targets from the stripe82 coadd
maskbits BOSS_TARGET1 16 QSO_KDE_COADD
# likelihood method
maskbits BOSS_TARGET1 17 QSO_LIKE

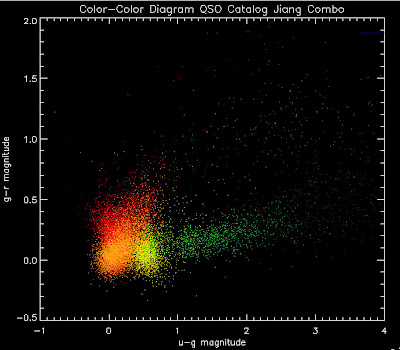
qsos = testqsosNow I need code to plot this histogram in color-color space with hotter colors in bins with higher likelihoods and cooler colors in bins with smaller likelihoods. Anyone have code that does this in IDL or python? Joe has some, but I'm having a hard time getting it to work.
b_u = 1.4
b_g = 0.9
b_r = 1.2
b_i = 1.8
b_z = 7.4
z_temp = qsos.Z_TOT
fu1 = qsos.PSFFLUX[0]
fg1 = qsos.PSFFLUX[1]
fr1 = qsos.PSFFLUX[2]
fi1 = qsos.PSFFLUX[3]
fz1 = qsos.PSFFLUX[4]
ivar_u1 = qsos.PSFFLUX_IVAR[0]
ivar_g1 = qsos.PSFFLUX_IVAR[1]
ivar_r1 = qsos.PSFFLUX_IVAR[2]
ivar_i1 = qsos.PSFFLUX_IVAR[3]
ivar_z1 = qsos.PSFFLUX_IVAR[4]
u1 = sdss_flux2mags(fu1, b_u)
g1 = sdss_flux2mags(fg1, b_g)
r1 = sdss_flux2mags(fr1, b_r)
i1 = sdss_flux2mags(fi1, b_i)
z1 = sdss_flux2mags(fz1, b_z)
sigu = sdss_ivar2magerr(ivar_u1, fu1, b_u)
sigg = sdss_ivar2magerr(ivar_g1, fg1, b_g)
sigr = sdss_ivar2magerr(ivar_r1, fr1, b_r)
sigi = sdss_ivar2magerr(ivar_i1, fi1, b_i)
sigz = sdss_ivar2magerr(ivar_z1, fz1, b_z)
ug = u1-g1
gr = g1-r1
ri = r1-i1
iz = i1-z1
icut = where(i1 GT 19.5)
nx = 50
ny = 50
imageQSO = fltarr(nx,ny)
imageAll = fltarr(nx,ny)
imageRatio = fltarr(nx,ny)
likeQSO = qsoqlike[icut]
likeAll = qsoslike[icut]
likeRatio = likeQSO/likeAll
qsoug = ug[icut]
qsogr = gr[icut]
populate_image, imageQSO, qsoug, qsogr, weight = likeQSO
populate_image, imageALL, qsoug, qsogr, weight = likeAll
populate_image, imageRatio, qsoug, qsogr, weight = likeRatio