I spent today trying to reproduce the result from this post when the reconstruction was working on the Mock Data.
I got it working... here is the code (also in ../logs/110509log.py):
#~~~~~~~~~~~~~~~~~
import numpy as N
from pylab import *
from matplotlib import *
import crlold as crl
from correlationData import *
import galsphere as gals
import datetime as dt
workingDir = 'run2010211_1354'
# Create file names
mockLRGfile,photofile,specfile,corr2dfile,corr3dfile,photo2dfile,photoCDfile,spec2dfile,\
spec3dfile,argumentFile,runConstantsFile,wpsMatrixFile,xiMatrixFile=makeFileNames(workingDir)
oversample = 5.
corrBins = 40.0 #should be one more than you want it to be
mincorr = 0.01
maxcorr = 20.0
convo = 180./pi
corrspace = (maxcorr - mincorr)/corrBins
corrarray = makecorrarray(mincorr, maxcorr, corrspace)
nbins = 40
boxside = 1000.0 #in Mpc/h
xicorrmin = 0.001
xicorrmax = 20.0
nxicorr = 40
#make redshift bins
#boxside = 1000.0 #in Mpc/h
rbinmin = 0 #in Mpc/h
rbinspace = boxside/40 #in Mpc/h
rbinmax = boxside/2.0 + 0.0001 #in Mpc/h
rbins = makeRbins(rbinmin,rbinmax,rbinspace)
nrbins = rbins.size - 1
photoCD = readPhotoCDfile(photoCDfile)
#Create a matrix to input the correlation function data
# Write wps Matrix to file
#The number of redshift bins you want to skip for the
#reconstruction (this removes noisy bins from calculation)
skip = 0
makeWPSmatrix2(workingDir,rbins,corrBins, wpsMatrixFile, skip)
#Create a matrix to input the correlation function data
# Write Xi Matrix to file
makeXimatrix2(workingDir,rbins,nxicorr,xiMatrixFile, skip)
#Plot wps correlation functions
numtoplot = int(nrbins-skip) #number of correlation functions you want to display on plot
numtoplot = 10
plotWPSScreen(skip, numtoplot, workingDir, rbins)
pylab.legend(loc=1)
#Plot xi correlation functions
plotXiScreen(skip, numtoplot, workingDir, rbins)
# compute the xi matrix using the spec sample,
npbins=40#the number of bins you want reconstructed
reconstructmax = 0.5 #in Gpc
reconstructionmin = 0.0 #in Gpc
phibins = linspace(start=reconstructionmin,stop=reconstructmax,num=npbins,endpoint=True)
#phibins =(linspace(2,npbins,num=npbins,endpoint=False)+0.5)*(1./npbins)
sximat=crl.getximat(th,rlos,phibins,rs,xispec,xierr)
# invert to recover phi
phircvd=crl.getCoeff(wps,covar,sximat)
for i in range(len(phircvd.x)):
if (phircvd.x[i] < 0):
phircvd.x[i]=0
print 'the recovered selection function is not normalized.'
outphi4 = phircvd.x
phibins4 = phibins
start = 1
end = npbins - 1
#junkbins = linspace(start=0,stop=0.5,num=100,endpoint=True)
junkbins = phibins
junk = histogram(photoCD/1000., bins=junkbins)
junkdata = junk[0]
plot(junkbins[start:end], junkdata[start:end], 'r.-', label = 'photometric data')
plot((phibins4[start:end]+1.0*phibsize), outphi4[start:end]*sum(junkdata[start:end])/sum(outphi4[start:end]), 'g-', label = 'phi')
plot(phibins4[start:end],outphi4[start:end]*sum(junkdata[start:end])/sum(outphi4[start:end]),'c.-',label='reconstruction %i'%npbins)
#plot((phibins4[2:]-1.5*phibsize)/1000, phi[2:]*size(photoCD), 'g-', label = 'phi')
xlabel('comoving distance (Gpc/h)')
ylabel('number of objects per bin')
title('Redshift Distribution')
from pylab import *
legend(loc=2)
#~~~~~~~~~~~~~~~~~
It involved reverting back to an old version of crl (I saved it as crlold.py) and an older version of the plotting.
Here are the correlation functions:
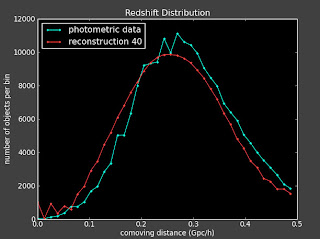
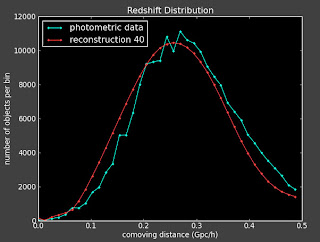
I am getting the same problems with slight changes to the binning, changing how well the reconstruction works. For instance the best reconstruction is here (with 40 bins):

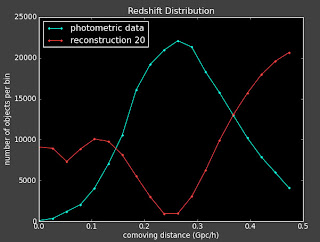
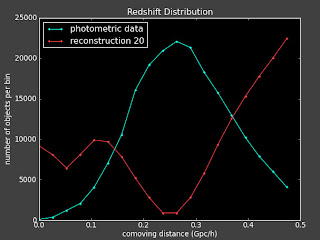
But if I change to 39 or 41 bins, I get a very different reconstruction:


Which makes me wonder if was just dumb luck that it worked for that particular binning in the first place.
I'm also running with a more interesting photometric distribution function to see how that looks. More to come....